The purpose of this device was to interact with real world objects. at MIT, in an attempt to help the disabled people produced a wearable gesture interface using a small camera and a projector. The work in this paper was especially inspired from the pioneering work of Mistry et al. Finally in Section 5, we conclude by presenting the simulation results obtained from the system exploring crystal anisotropy of silicon during its nanoindentation. In Section 4, the nanoindentation study using the VAMDS system is described. Section 3 presents our proposed VAMDS methodology. Section 2 briefly reviews earlier efforts in design and simulation systems driven by haptic devices. We demonstrate our approach and describe our vision-augmented molecular dynamics simulation (VAMDS) system on the nanoindentation of a silicon crystal silicon substrate.
Jmol simulation simulator#
This approach progresses the current state of the art via a step change in understanding of the complex interactions as well as allowing the rapid design of novel molecular structures using an intuitive way of interaction with a simulator by means of hand gestures.
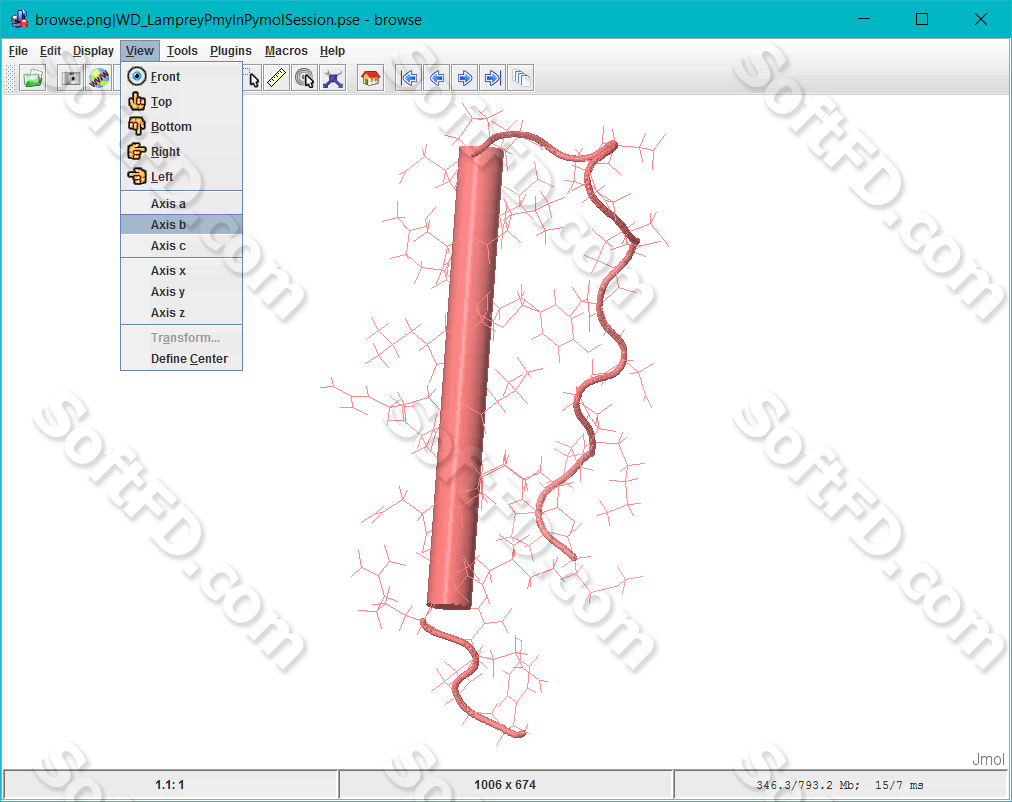
Our approach entails the use of computer vision and specifically gesture recognition to enable the user to interact with the simulated system exploiting the numerous degrees of freedom available with hand gestures in 3D space, breaking away from the constraints of traditional methods of interaction involving a high level language based inputs which became the motivation for this paper. This research is focused on the development of a virtual environment which offers an entirely new way to explore complex energy landscapes by allowing the user to directly interact with the simulated molecular system. These approaches have a high computational cost as they tend to get confined within local regions of the energy landscape spending many CPU during a simulation run. IntroductionĬomputational simulation approaches, such as molecular dynamics (MD), Monte Carlo, or coarse grained MD, are widely employed by chemists and materials scientists to predict and understand the behaviour of materials. We anticipate that our proposed system will open up new horizons to the current methods on how an MD simulation is designed and executed. A long-range (Screened bond order) Tersoff potential energy function was used during the simulation which revealed the value of hardness and elastic modulus being similar to what has been found previously from the experiments. The proposed system was tested by simulating the crystal anisotropy of crystalline silicon during nanoindentation. The end result is that users with limited expertise in developing molecular structures can now do so easily and intuitively by the use of body gestures to interact with the simulator to study the system in question.
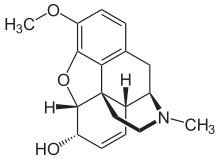
The hand gestures are used to pick and place atoms on the screen allowing thereby the ease of carrying out molecular dynamics simulation in a more efficient way. The system is novel in its concept as it enables the user to directly manipulate the atomic structures on the screen, in 3D space using hand gestures, allowing the exploration and visualisation of molecular interactions at different relative conformations. We present a user-friendly vision-augmented technique to carry out atomic simulation using hand gestures.
